Suppose you're a machine-learning researcher trying to build a model that could help plan for the COVID-19 pandemic. You want to incorporate a disease simulator into the model, but it's written in the C++ programming language, rather than an existing machine-learning workflow like PyTorch or TensorFlow.
Traditionally, this means you would have to spend more time learning C++ and rewriting it in the other framework than you spend actually building a model to solve the problem.
A team from MIT CSAIL recently developed a clever work-around: a compiler plug-in called Enzyme that allows you to import arbitrary foreign code into systems like TensorFlow without having to rewrite it.
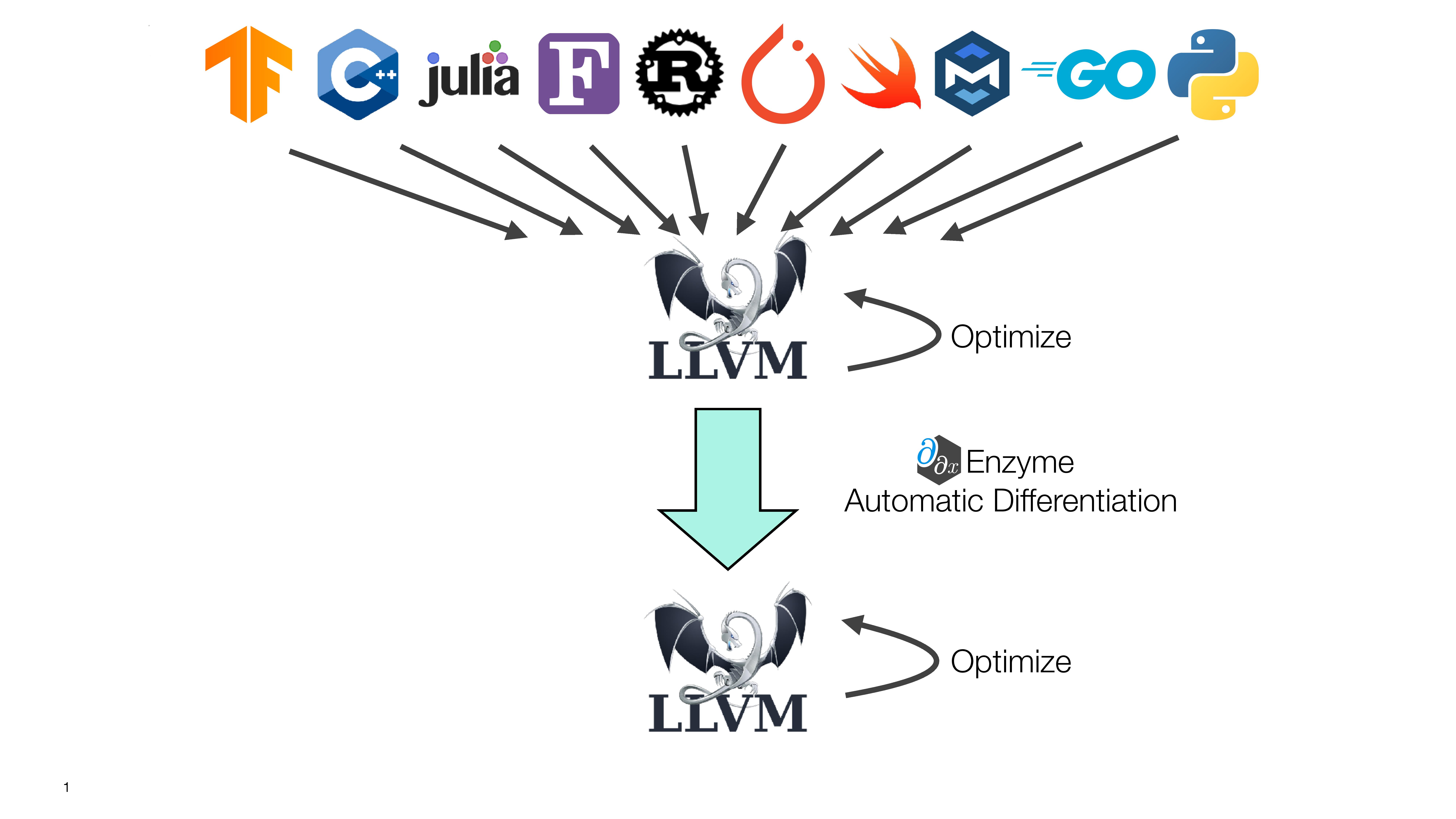
The researchers say that Enzyme reduces the amount of work needed to apply machine learning to new domains, and is compatible with virtually any programming language that uses the LLVM framework, including C, C++, Rust, Swift and Julia (which originated at MIT).
“We hope Enzyme will help advance a range of scientific disciplines and bridge the gap between the machine learning and scientific computing communities,” says PhD student William S. Moses, lead author of a new paper about the system. “By allowing these groups to share tools and more easily interoperate, Enzyme may allow for improved approaches in many fields, from public-health analysis to climate simulation.”
Enzyme solves a specific problem with respect to “automatic differentiation” (AD), a family of algorithms that take derivatives of functions implemented in computer programs. AD is used within fundamental machine learning practices like backpropagation, as well as many areas of scientific computing.
It has historically been assumed that a high-performance tool like AD is only practical at the highest language level, but Moses says that Enzyme shows that it’s possible at lower levels. In fact, working at a lower level enables the system to run optimizations before differentiation, which allows it to execute programs up to four times faster than existing tools.
In the field of scientific computing, Enzyme also takes out a lot of the manual work that programmers loathe. For example, if you’re developing a simulator for a physics engine, Enzyme could be used to calculate derivatives that previously had to be done by hand.
The team has been excited to see Enzyme already receive some interest from the global programming community. As next steps, the team plans to improve Enzyme’s interface and improve the plug-in so that it can instantly enable programmers to import external codebases into their workflows.
The team will be presenting the project this week as a spotlight talk at the annual Conference on Neural Information Processing Systems (NeurIPS).
The project was supported in part by the Defense Advanced Research Projects Agency (DARPA), the Department of Energy, Los Alamos National Laboratories, the National Science Foundation, and the United States Air Force Research Laboratory.